Medical Segmentation Decathlon
Generalisable 3D Semantic Segmentation
Notifications
- The Decathlon challenge paper is now on Nature Communications and ArXiv
- The Decathlon dataset specific paper is also on ArXiv
- New rolling competition and leaderboard is now available here
Aim
With recent advances in machine learning, semantic segmentation algorithms are becoming increasingly general purpose and translatable to unseen tasks. Many key algorithmic advances in the field of medical imaging are commonly validated on a small number of tasks, limiting our understanding of the generalisability of the proposed contributions. A model which works out-of-the-box on many tasks, in the spirit of AutoML, would have a tremendous impact on healthcare. The field of medical imaging is also missing a fully open source and comprehensive benchmark for general purpose algorithmic validation and testing covering a large span of challenges, such as: small data, unbalanced labels, large-ranging object scales, multi-class labels, and multimodal imaging, etc. This challenge and dataset aims to provide such resource thorugh the open sourcing of large medical imaging datasets on several highly different tasks, and by standardising the analysis and validation process.

Data
All data will be made available online with a permissive copyright-license (CC-BY-SA 4.0), allowing for data to be shared, distributed and improved upon. All data has been labeled and verified by an expert human rater, and with the best effort to mimic the accuracy required for clinical use. To cite this data, please refer to https://arxiv.org/abs/1902.09063
Schedule & Guidelines
The MSD challenge tests the generalisability of machine learning algorithms when applied to 10 different semantic segmentation tasks. The aim is to develop an algorithm or learning system that can solve each task, separateley, without human interaction. This can be acheived through the use of a single learner, an ensable of multiple learners, architecture search, curriculum learning, or any other technique, as long as task-specific model parameters are not human-defined.
Participants are expected to download the data, develop a general purpose learning algorithm, train the algorithm on each task training data independently without human interaction (no task-specific manual parameter settings), run the learned model on the test data, and submit the segmentation results.
Tasks
Liver Tumours
Target: Liver and tumour
Modality: Portal venous phase CT
Size: 201 3D volumes (131 Training + 70 Testing)
Source: IRCAD Hôpitaux Universitaires
Challenge: Label unbalance with a large (liver) and small (tumour) target

Brain Tumours
Target: Gliomas segmentation necrotic/active tumour and oedema
Modality: Multimodal multisite MRI data (FLAIR, T1w, T1gd,T2w)
Size: 750 4D volumes (484 Training + 266 Testing)
Source: BRATS 2016 and 2017 datasets.
Challenge: Complex and heterogeneously-located targets
Hippocampus
Target: Hippocampus head and body
Modality: Mono-modal MRI
Size: 394 3D volumes (263 Training + 131 Testing)
Source: Vanderbilt University Medical Center
Challenge: Segmenting two neighbouring small structures with high precision
Lung Tumours
Target: Lung and tumours
Modality: CT
Size: 96 3D volumes (64 Training + 32 Testing)
Source: The Cancer Imaging Archive
Challenge: Segmentation of a small target (cancer) in a large image

Prostate
Target: Prostate central gland and peripheral zone
Modality: Multimodal MR (T2, ADC)
Size: 48 4D volumes (32 Training + 16 Testing)
Source: Radboud University, Nijmegen Medical Centre
Challenge: Segmenting two adjoint regions with large inter-subject variations
Cardiac
Target: Left Atrium
Modality: Mono-modal MRI
Size: 30 3D volumes (20 Training + 10 Testing)
Source: King’s College London
Challenge: Small training dataset with large variability
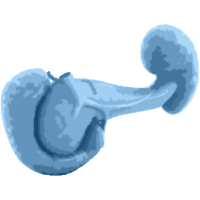
Pancreas Tumour
Target: Liver and tumour
Modality: Portal venous phase CT
Size: 420 3D volumes (282 Training +139 Testing)
Source: Memorial Sloan Kettering Cancer Center
Challenge: Label unbalance with large (background), medium (pancreas) and small (tumour) structures.
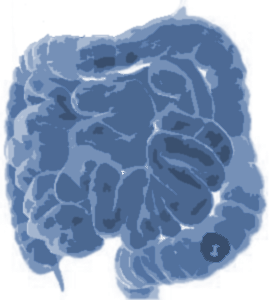
Colon Cancer
Target: Colon Cancer Primaries
Modality: CT
Size: 190 3D volumes (126 Training + 64 Testing)
Source: Memorial Sloan Kettering Cancer Center
Challenge: Heterogeneous appearance
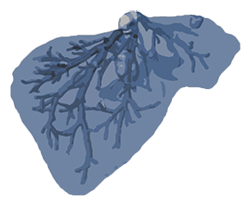
Hepatic Vessels
Target: Hepatic vessels and tumour
Modality: CT
Size: 443 3D volumes (303 Training + 140 Testing)
Source: Memorial Sloan Kettering Cancer Center
Challenge: Tubular small structures next to heterogeneous tumour.

Spleen
Target: Spleen
Modality: CT
Size: 61 3D volumes (41 Training + 20 Testing)
Source: Memorial Sloan Kettering Cancer Center
Challenge: Large ranging foreground size
Assessment Criteria
The challenge aims to optimise algorithms for generalisability and not necessarily attempting to achieve state-of-the-art performance on all 10 tasks. Thus, a small subset of classical semantic segmentation metrics, in this case the Dice Score (DSC) and a Normalised Surface Distance (NSD), will be used to assess different aspects of the performance of each task and region of interest. These metrics are implemented here. These different metrics will be combined to provide a single task performance metric, and then meta-analysed to obtain an overall performance metric. Further details on the statistical analysis methodology is described here.
Due to the complexity of the challenge, the metrics (DSC,NSD) were chosen purely due to their well-known behaviour, their popularity, and their rank stability [see https://arxiv.org/abs/1806.02051]. Having simple and rank-stable metrics also allows the statistical comparison between methods. Note that the proposed metrics are not task-specific and not task-optimal, and thus, they do not fulfill the necessary clinical criteria for algorithmic validation of each independent task.
Organising Team and Data Contributors

M. Jorge
Cardoso
King's College London
Role: Leadership, Conceptual Design, Data Pre-Processing, Stats and Metrics Committee

Amber
Simpson
Memorial Sloan Kettering
Cancer Center
Role: Data Donation (Pancreas, Colon Cancer, Hepatic Vessels, Spleen), Conceptual Design

Olaf Ronneberger
Google Deepmind
Role: Conceptual Design, Metrics Committee

Bjoern
Menze
Technische Universität München
Role: Data Donation (Brain Tumours, Liver Tumours), Stats and Metrics Committee

Bram
van Ginneken
Radboud University Medical Center
Role: Data Donation (Prostate), Conceptual Design

Bennett
Landman
Vanderbilt University
Role: Data Donation (Hippocampus), Conceptual Design, Metrics Committee

Geert
Litjens
Radboud University Medical Center
Role: Data Donation (Prostate)

Keyvan
Farahani
National Institutes of Health
Role: Data Donation (Lung Tumours)

Ronald
Summers
National Institutes of Health Clinical Center
Role: Conceptual Design

Lena
Maier-Hein
DKFZ German Cancer Research Center
Role: Conceptual Design, Statistical Analysis Committee

Annette
Kopp-Schneider
DKFZ German Cancer Research Center
Role: Conceptual Design, Statistical Analysis Committee

Spyridon
Bakas
CBICA, University of Pennsylvania
Role: Data Donation (Brain Tumours)

Michela
Antonelli
University College London
Role: Conceptual Design, Data Preparation, Validation System Setup, Challenge Day-to-day Support
Frequently Asked Questions
Bellow are some of the most asqued questions received over email or other contact media.
Who should I contact if I have any questions?
You should contact the organisers by emailing medicaldecathlon@gmail.com
Can I change the learning rate or any other parameter (regularisation, depth) per task?
The ultimate aim of the challenge is for the winning algorithm to “just work” on any future unseen task, without requiring any extra inputs or human interaction. Thus, the learning algorithm/script/parameters cannot be changed manually per task. Parameters can only be made task specific if the parameter is found automatically within the learning system itself (e.g. through cross-validation).
Is it permitted to train with additional datasets that are not part of the challenge?
Using task-specific external data to pre-train the nework would be considered a task-specific change and is thus not allowed. You can use a general purpose pre-trained network to initialise all the tasks, but not a pre-trained network which is task specific. In short, you can do anything methodologically as long as it is not task-specific and would work on any future new task.
Are we supposed to submit the segmentation masks or the code to the challenge?
Only segmentation masks are required. We incentivise that people make the code available, but this is not a requirement.
Is the validation process handled as a black box, such as a virtual machine?
Yes, a docker container that will do all the validation on the server. We will release the validation scripts for public scrutiny, but not the validation labels.
Do I need to identify myself when submitting results?
There are not requirements of identifiability for submissions. You will however have to identify the participants names and institutions if you want to be part of a future journal submission describing the outcomes of the challenge.
Sponsorship

